The last time I played with Google Street View, I saw my car in the driveway and could practically count the teeth on a jack o’ lantern on my porch. I thought, wouldn’t it be amazing if I could drop-land on the surface of a nanoparticle and take a look around the scenery? What about on any other surface for that matter? Your kitchen table or the palm of your hand? What does it look like over there—what molecules are stuck to the ground; are there rolling hills or flat vistas? Even when something looks simply flat to our eyes, the true nature of the surface is usually far from simple.
Discovering what surfaces look like is a notoriously difficult task for scientists. One reason surfaces matter is because the features and fate of nanoparticles are greatly determined by their surfaces. When a particle floats through the bloodstream or settles into the flora of a lake, the cells and plants “see” the surface of the particle. They will interact with the particle differently whether the surface is highly charged or neutral, pillowy soft or jagged and rough. We often anchor small organic molecules or polymers to particles in order to govern their interactions with the environment. For example, we can wrap a particle with the cushy polymer poly(ethylene glycol) (PEG) to thwart the attachment of “sticky” proteins. Biomedical researchers often do this when they want the surface of their technology to remain unfouled by the various things floating around in our bodies.
However, proof of the nature of the surface can elude us: interfaces (like between a solid and a liquid) are very challenging to study. Rather than use one tool, we collect a dossier of data that hints at its attributes. X-ray photoelectron spectroscopy tells us which atoms are present, atomic force microscopy probes the roughness of the surface, and zeta potential measurements spit out the surface charge. But sometimes we want harder evidence of the tethered organic molecules present.
Although understanding an interface is difficult, there’s a gold standard method for revealing the structure of pure organic molecules like pharmaceutical drugs: nuclear magnetic resonance spectroscopy, or NMR. If you’ve ever had an MRI scan of your brain or other organs, you are intimately acquainted with the workings of NMR. MRI’s more formal moniker, Nuclear Magnetic Resonance Imaging (NMRI), was cleverly abridged to spare the patient the alarming “nuclear” term. But despite conjuring images of bombs and radiation, “nuclear” is not a scary word at all: Every atom has a nucleus, so we’re all full of nuclei.
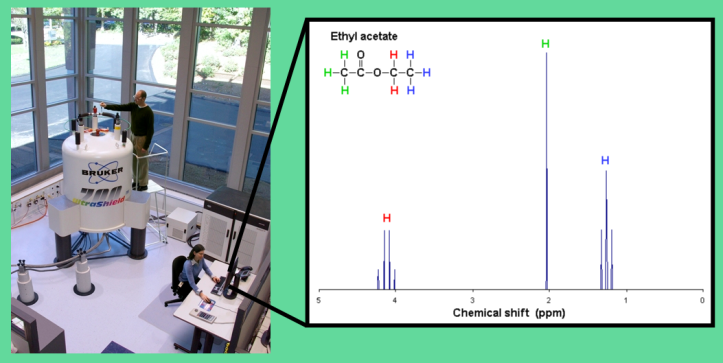
Some consider NMR a black box of magic: Insert a tube of your molecule into a giant cosmic-looking cylinder, which spits out a molecular fingerprint in the form of a spectrum.
A trained chemist can interpret the peaks—sharp or broad, spaced out or squeezed together—to figure out or confirm the molecule’s structure.
That hulk of a machine is actually a huge magnet, which encases the molecules in a strong magnetic field. (On more than one occasion, I’ve erased credit cards or felt a strong magnetic tug on my earrings as a result of getting too close to it.) Conveniently, some atomic nuclei are magnetic. A magnetic nucleus is a miniscule version of an ordinary refrigerator magnet, so it interacts with the magnetic field applied by the machine. NMR teases the nuclei in a molecule with electromagnetic energy, and we can learn a lot about those nuclei by seeing how they respond.
I find an analogy is helpful at this point. Consider a field of hungry wolves. Starving and lost, they run randomly in every possible direction, in search of prey. The alpha male howls from the edge of the field, and the majority of the pack obediently turns to face him. But, suddenly, a bolt of lightning strikes the opposite end, and the riled-up wolves whip toward the sound. They yelp and bark a bit. The alpha male, not about to be overshadowed, howls again, and slowly the wolves turn their attention back to him. The hungrier ones take a little longer to settle down.
In this analogy, the wolves are our atomic nuclei and the howling alpha male is the magnetic field of the NMR spectrometer. When we submerge our molecule into the NMR magnet, the randomly-oriented nuclei align parallel to the magnetic field. In order to elicit an NMR signal from the nuclei, we inject them with energy in the form of radio waves—the equivalent of rousing the wolves with lightning. The extra energy encourages the nuclei to temporarily turn against the field. However, after the pulse of energy is over, the nuclei slowly “relax” back parallel to the field.
This relaxation process is diagnostic for every nucleus, dependent on its location in the molecule and nearby atoms. As the nuclei relax, they emit “resonance” signals, sort of like the wolves’ yapping. The NMR machine collects the resonance signals from all the nuclei in the molecule, and then does a bunch of math to transform them into a unique spectrum of peaks, a “map” of the atoms in the molecule.
NMR of nanoparticles has the capacity to decipher many of the field’s current mysteries. The nuclei maps may be able to expose the general topography and character of the particle surface: Is it a sparse landscape of a single type of molecule, like a desert of Joshua trees? Or is it densely packed with various molecules, like the mix of spruce, pines, and firs of the Appalachian forests? Do the molecules bend and flap around freely, like those trees in the wind, or are they rooted and unyielding? Not unlike Google Maps after all.
… We wish. In reality, NMR of particles is currently more like the early beta versions of MapQuest. It can provide a fuzzy idea of the particle landscape, but nowhere near the level of complex detail required to really understand the surface. Why not?
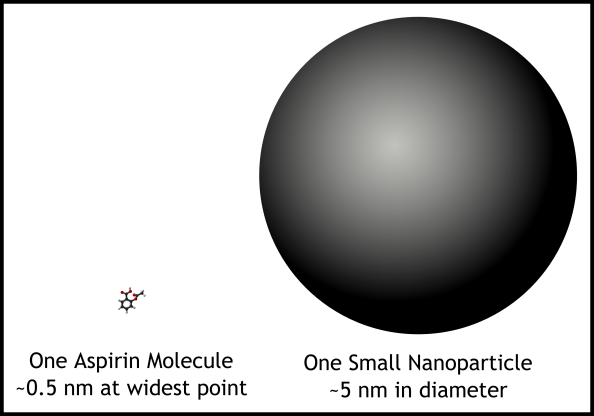
On numerous other posts on this blog (How to Avoid Bitter Coffee…, Why are Nanomaterials so special?, Nature’s Nanomachines Harnessed to Enable Healthier Delicious Ice Cream) we’ve hammered in the point that nanoparticles are tiny. However, they’re actually huge when compared to most organic molecules, like those that make up medications like aspirin and ibuprofen. NMR is optimized for small, agile molecules like these, not slow, bulky nanoparticles.
For a tiny nuclear magnet to emit a clear resonance signal, it must be able to tumble around freely and nimbly. When it’s anchored to something as “heavy” as a nanoparticle, its movement is restricted—like a wolf whose paws are sinking into mud during a storm. This restriction dramatically affects the nucleus’s relaxation process and muffles its resonance signals (as the rain drowns out the wolf’s bark), so the resulting spectrum is much harder to interpret.
NMR is also not a very sensitive technique, so it can’t detect molecules below a certain concentration. Many of our nanoparticle syntheses give us scant amounts of product—often way below this detection limit.
If NMR provides its own set of challenges, why do we bother with it at all? Under the right conditions, NMR can reveal things that other techniques can’t. X-ray photoelectron spectroscopy or Fourier transform infrared spectroscopy, for example, might simply disclose the atoms or bonds present, whereas NMR can depict a complete portrait of the molecular structure. So, we pool our arsenal of data from a wide variety of techniques to create the most complete picture possible of the nanoparticle surface. Maybe someday we’ll have our nanoparticle version of Google Street View.

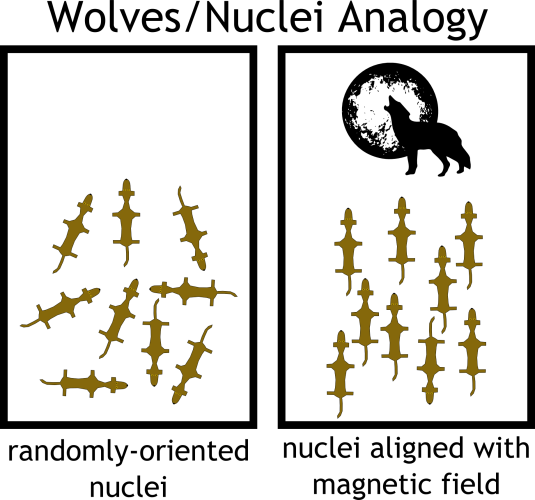

[…] a nanomaterial once it’s been in contact with a cell membrane? Use nuclear magnetic resonance (blog entry), transmission electron microscopy, X-ray photoelectron spectroscopy, or atomic force microscopy. […]