Not too long ago, a team of frustrated scientists glanced up from their computer screens and saw the untapped potential beyond their lab doors. They had spent years trying to unravel the mysteries of protein folding, but some just couldn’t seem to be solved. Why not harness the brainpower of the public? The biochemists disguised their puzzles as a colorful video game, called Foldit, and watched with amazement as the living-room researchers cracked puzzles that had stumped even their cutting-edge computers.
“Citizen science” is now in vogue, as scientists are realizing that even those without science backgrounds can make huge discoveries. This is especially true in fields limited by computational power, time, or the sheer number of experiments to try, like protein folding. After all, humans have intuition and creativity (where computers do not) and can work en masse (where academic labs are limited).
Nanotechnology is entering a critical stage as scientists puzzle over how to make tiny, three-dimensional machines that are functional and safe. We have an overwhelming number of things to test out, but not enough time. Can nanoscientists enlist the collective intelligence of living-room scientists to solve these problems, just as protein scientists have done? I think the answer is yes!
First, to the story of how the citizen biochemists helped crack protein folding.
Proteins are the workhorses of life within our cells. A young protein begins its life as a floppy, formless string. It’s helpless to do anything until it folds up into the unique three-dimensional shape that is its functional, “adult” form. Coming-of-age is turbulent, and the adolescent protein experiments with its persona by folding into different shapes, some high in energy and defiant, others low-energy and relaxed. When it makes mistakes, the guiding laws of physics gently steer the protein back on track. Finally, the protein settles happily into its functional form, known to scientists as the native form—usually the lowest-energy structure. Most proteins fold into the correct shape, but in rare instances a rebellious protein will go too far down the wrong path and end up unrecognizable. This misfolded protein can be a bad influence on healthy proteins, forming clumpy, dysfunctional gangs that rule the dark underbelly of human health and affect some very grim illnesses—neurodegenerative diseases like Alzheimer’s, Parkinson’s, or mad cow, for example. Scientists are desperate to eradicate the rebels, but they first need to know what makes them tick: What does a well-folded protein look like, and what goes wrong on the way there?

To answer these questions, scientists visualize a protein’s rocky adolescence as a 3-D energy landscape, which looks like a snapshot of the Himalayas. Every spot on this landscape corresponds to a potential shape the protein could fold into, with jagged peaks reflecting the high-energy shapes and dramatic craters signifying the low-energy ones. At the bottom of a specific low-energy hole awaits the native structure, and it’s the protein’s journey to this hole that marks the enigmatic protein folding process. Scientists use state-of-the-art computer software to discover the protein’s path and final shape. The protein is dragged through an endless puberty along the energy landscape, pried apart and contorted into grotesque positions. The computer scrutinizes the energy of each position, adjusts the protein slightly, and repeats this unhappy ritual until the protein settles into a very low-energy structure, marked by a dint in the energy landscape. “Native state!” the computer declares, and stops there.

But it’s wrong. The landscape is difficult to navigate without a map, and although that dint may look deep from up close, there might be a deeper one just over that mountain… The computer, clueless to the location of the real native state and constrained to a rigid set of algorithms, won’t risk trekking back up the hill in search of an unknown possibility. It gets stuck.
That’s where citizen science comes in. Our intuitive understandings of big picture goals and manipulating 3-D space give us humans a significant edge over computers. Whereas the computer was tricked by that shallow hole, a person may evaluate the protein’s structure, remember the last time this happened, and imagine better possibilities. The Foldit interface includes an exploration map that mirrors the energy landscape—there are regions of high and low scores that other players have unearthed, as well as uncharted territory, structures that have not been tested. Humans are much more likely than computers to dig around these frontiers, risking momentary low scores for the ultimate gold. In one astonishing case, Foldit players figured out the structure of a monkey AIDS virus protein, which had stumped computers for ages; their correct model will help scientists design new drugs. The gamers’ pseudonyms (“Contenders” and “Void Crushers”) appear next to the scientists’ in the ensuing scientific paper, and the Foldit creators are optimistic that the game will continue to solve many of the protein folding puzzles that still remain.
So, where’s the scientific gold for citizen nanoscientists?
Many of the million-dollar questions of nanoscience parallel the mysteries of protein folding. An oft-touted promise of nanotechnology is our ability to “program” individual particles to do whatever we want. The tiny particles are born in a lab with their fates sealed. Scientists program the particles to stick together by tethering choosy “recognition” molecules to their surfaces. In what’s known as self-assembly or bottom-up design, the nanosize building blocks scramble around and ultimately organize themselves into an assortment of larger shapes—spheres, tubes, and, the eventual hope is, nanoscale machinery. These may range from cellular big rigs that deliver drug cargo and tow away deadly bacteria, to life-size screws and nails that grow artificial scabs to heal themselves when cracked… because they are constructed of nanoparticles that know what to do.
If this sounds very sci-fi, try your hand at self-assembly. Imagine that you’ve got a bucket of blocks, all sorts of shapes and sizes. These are your “nanoparticles.” When you shake the bucket, I want all the yellow spheres to stick together, and all the blue cubes to stick together. “No sweat,” you say, brushing your shoulder, and put Velcro strips on all the yellow spheres and magnets on the blue cubes. You shake, and it works—that’s self-assembly! But that was just a warm-up. Now, build this 3-D scene: On one face of a green rectangle, spell out “wonder” using small spheres in rainbow order. Underneath, I want a replica of the Great Pyramids. On the middle pyramid, in the exact center of the face farthest from “wonder,” construct a crude Sphinx out of yellow ovals and cubes. Give it blue eyes, and rotate its entire body 45˚ clockwise from the base of the pyramid. Remember, this scene must appear as if by magic when you shake the bucket, so pick and place your recognition units wisely.
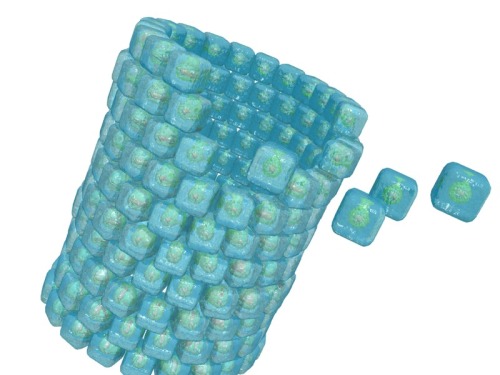

You can see that self-assembly gets complicated quickly. You’d need more “sticky” things than Velcro and magnets, and would have to cut out tiny pieces of each and place them in very specific patches on the blocks. With the right tools and lots of patience, though, you could probably build ancient Egypt. Working on the nanoscale makes it that much harder. For fine control over nanoparticles, scientists need sophisticated types of recognition molecules. They first design a patterned particle blueprint using a mix of intuition and computation, then spend months or years trying to make the actual particle, which often fails to assemble itself as predicted. (Maybe the Sphinx’s eyes appeared on its back, or “wonder” came out as “mother”.) Back they go to the drawing board. The whole process is agonizingly slow.
So, let us call on the citizen scientists. Earlier, you tried your hand at self-assembly using Velcro’ed blocks. But how easy would it be for casual gamers to program their own particles using molecules, if even computer-enabled chemists struggle with it? These tiny molecules serve as nanoscopic Velcro, and can be attached to nanoparticle surfaces to control the way each particle behaves. Although the interactions between molecules can be chemically complex, they can be distilled to a simple set of rules. These rules should feel natural to most people, because we’re familiar with them, just on a macroscopic scale. Take shape complementarity. Only your teeth fit in your retainer; only that one piece fits in the corner of the jigsaw puzzle. Likewise, molecules come in a variety of shapes, from rigid zig-zags to hollowed-out pumpkins—and matching geometries fit together. Or another example: we know well that oils don’t mesh with water; pour bacon grease down the sink and you’ll never forget the fat glob that forms. Hydrophobic molecules like those in oils are drawn to each other by shared distaste for water, huddling together like a middle school clique to avoid contact with their common enemy. This effect alone drives a huge share of self-assembly processes.
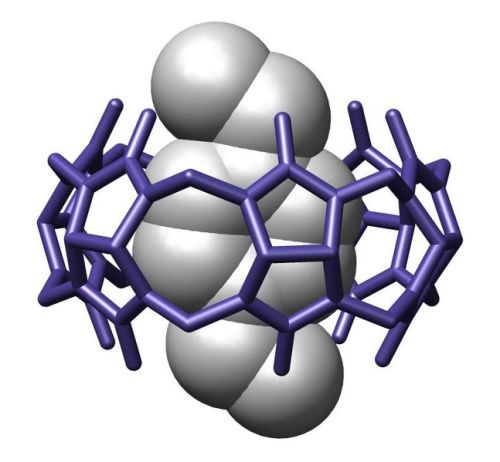
Molecules don’t need to be baffling. The ideas underlying self-assembly and molecular recognition are intuitive to people—and easy to package in user-friendly tools and cartoons. Game players could decorate their particles’ surfaces with recognition units, and let them loose to see if they self-organize into the desired architecture. Points would be awarded not only to well-programmed particles, but also to those that assemble into the most innovative, complex architectures. Nanoscale factories with conveyor belts to sort and package drugs within the body? Zoos of molecular animals that could use their hinged jaws to eat and ball-in-socket joints to run? Why be limited to what we know? After all, the breakneck pace of nanotechnology requires that we reward the pioneers, and humans are certainly better than computers at pushing the frontier.
Author’s note: Want to try your hand at citizen science? Here’s a list of some existing citizen science projects for you to check out. If you have one to share, let us know in the comments!
* Foldit: Fold proteins
* EyeWire: Map the brain
* EteRNA: Design RNA molecules
* Stardust@home: Find interstellar dust
* Galaxy Zoo: Classify galaxies
* SETI@home: Search for aliens
* BudBurst: Monitor climate change
* eBird: Track birds
